UPDATE: Additional simulations by Andrew Millard using different recombination rates for men and women indicate that half-siblings and aunts/uncles/nieces/nephews can sometimes be distinguished from one another if they are on one’s paternal side but not if they are on the maternal side. Please bear that in mind as you read this post.
With the exception of identical twins or parents and their children, a given relationship has a range of possible shared DNA. Grandparents and their grandchildren can share from about 1156 cM to 2311 cM, with an average of 1766 cM. (The values in this post were taken from user-reported values in the 2017 version of the Shared cM Project, by Blaine Bettinger; “cM” stands for centimorgan, the unit we use to quantify shared DNA in genetic genealogy.)
(A note for newbies: Words in bold in this post are defined in the glossary.)
Most family relationships have ranges that overlap with at least one other relationship, meaning that there’s no way to tell the possibilities apart using only the amount of shared DNA. For example, the reported range for aunt/uncle and niece/nephew (an avuncular or nibling relationship) is 1349–2175 cM (average 1750 cM) and for half siblings it’s 1317–2312 cM (average 1783 cM). Compare to 1270–2365 cM (average 1766 cM) for grandparents. In an unknown parentage search or when faced with an unexpected match, we often want to know as much as possible before approaching the other person. Sometimes non-genetic factors can help to determine the relationship—if a DNA match is only 15 years older, they cannot be a grandparent—but often there is no way to tell.
Or, is there?
The Same Isn’t Always the Same
Although they have nearly identical averages and ranges of shared DNA, the grandparent–grandchild, avuncular, and half-sibling relationships differ fundamentally in how the DNA is inherited. Let’s take a look. Before we do, I’ll remind you that we each have two copies of each autosomal chromosome—one inherited from each parent—and each copy is a mixture of the two chromosomes of that parent’s parents (our grandparents). The mixtures occur when gametes (eggs and sperm) are formed, during a process called crossing over.
Grandparent Versus Grandchild
I am going to walk you through simplified examples for the three relationships, all using chromosome 1 with two crossovers per generation. Of course, in the real world, we have 22 autosomal chromosome pairs, any of which can cross over one or more times or not at all.
The first example compares a grandfather to his grandchild (his daughter’s child).
Grandpa’s two copies of chromosome 1 cross over where the black X-like marks are, and one recombined chromosome is inherited by his daughter. She, in turn, has two copies of chromosome 1 (the other came from her mother) that recombine, and the child inherits one of them. (The grandmother’s chromosomes also recombine before being passed to the mother, but because they are not relevant to the grandfather–grandchild comparison, her chromosome 1 is shown in black for now.) A total of four crossovers have taken place, and the grandfather and grandchild match along a single, rather large segment of DNA (yellow bar). Note that a crossover took place within that segment, but with the tests we use for genetic genealogy, all we’ll see is a single large matching segment.
Half Sibling Comparison
Now consider half siblings who share a mother. After recombination, her first child receives a copy of chromosome 1 that looks like this:
And her second child receives a copy of chromosome 1 that looks like this:
Comparing the two half siblings to one another, we can see where they match one another (the yellow bars).
In both the grandparent–grandchild scenario and the half-sibling one, there are two generational steps (grandparent-to-mother, mother-to-child and mother-to-half-sib-A, mother-to-half-sib-B, respectively). What’s more, I used the exact same crossover points in the two examples. The grandpa’s two crossover points in the first example are exactly the same as the ones that produce Half-Sibling A in the second example, and the mom’s two crossover points in the first case are exactly the same as those leading to Half-Sibling B in the second.
But we end up with completely different patterns of sharing! One large segment shared between grandpa and grandchild versus two smaller segments shared between half siblings.
That’s gotta make you wonder: Can we possibly tell a grandparent from a half sibling using DNA alone if we consider not just how much DNA is shared but also how many segments are involved?
Avuncular Relationships
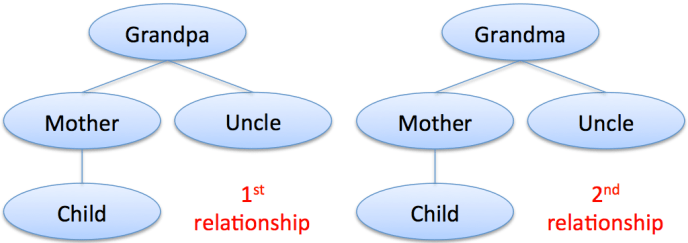
Before we answer that question, let’s look at the third case with an overlapping range, the avuncular relationship. This one is a bit more complicated to depict, for two reasons. First, there’s an additional generational step involved, and second, it’s actually a double relationship.
What I mean by “double relationship” is that the child and uncle are related once through the child’s grandfather and again through the grandmother.
(If they were only related through one grandparent, the uncle would be a half uncle.)
In this example, I will use the grandfather–grandchild example from above, but instead of using a generic black chromosome for Grandma, we’re going to work her into the equation, too.
Let’s divvy up grandma’s chromosomes into her two children, the child’s mother and the uncle. Again, I’m using two crossovers per chromosome pair and placing them whimsically. The grandmother’s two chromosomes recombine into the uncle like this:
And into the mom like so:
Now, let’s take the mom’s paternal chromosome from the very first example above and replace the black stand-in chromosome with the maternal chromosome that we just created. The crossover points are the same as in that first example. Now, the recombination that produces the child looks like:
In this case, the child’s maternal chromosome 1 is comprised of segments that originally came from both grandparents.
Finally, let’s compare the child to the uncle to see where they share DNA segments (the yellow bars).
Interesting! We used the same number of crossovers per generation, and we’re looking at three relationships that would be indistinguishable based solely on total amount of shared DNA, but we see different patterns when we look at the number of segments.
How can we apply this to genetic genealogy?
Simulations by Dr. Andrew Millard
It turns out that the idea that there might be differences among these relationships isn’t new. Scientists from 23andMe published a scientific paper entitled “Cryptic distant relatives are common in both isolated and cosmopolitan genetic samples” back in 2012. They include a figure that indicates almost no overlap between grandparent–grandchild (the orange “cloud” of dots) and avuncular (the yellow cloud) relationships when both total shared centimorgans and the number of segments are considered. However, this paper didn’t include half siblings.

In 2013, William Hill and Ian White of the University of Edinburgh, UK, did a study that included half siblings, but they counted segments far smaller than we use in genetic genealogy (down to 1 cM) and reported their results in awkward tables.
To counter these limitations, Dr Andrew Millard of the Durham University, UK, designed a computer simulation that mimics crossovers, segment inheritance, and the matching parameters used by GEDmatch. He then simulated thousands of avuncular (UN in the chart below; the black cloud), half sibling (HS; the red cloud), and grandparent–grandchild (GP; the blue cloud) relationships and plotted each simulated result by the total amount of shared DNA and the number of segments.

Here’s another view, with the red and black colors reversed to give you a different perspective on the overlap area.
Yes! We can often tell these relationships apart! The grandparent–grandchild relationship is almost always distinct from the other two, and while there is a fair amount of overlap between the avuncular and the half-sibling clouds, there are also regions where one or the other is the only possibility or is significantly more likely.
Try It!
To use this chart, do a one-to-one comparison of two kits at GEDmatch. At the bottom of the resulting table, find the Total of segments > 7 cM and the number of matching segments, like shown.
Then find the place on the plot that corresponds to the total shared cM (the left–right axis) and the number of shared segments (the up–down axis) and observe which “cloud” is densest. Let’s do a few examples.
Example 1: Two people at GEDmatch share 1530 cM across 26 segments. The yellow ❶ on the example plot shows where that match falls. That’s firmly in the blue (grandparent/grandchild) cloud with neither of the other two relationships having that few segments, so they are grandparent and grandchild to one another.
Example 2: Two people at GEDmatch share 1756 cM across 51 segments. The yellow ❷ on the example plot is in a dense part of the black (avuncular) cloud and in a very sparse part of the red (half sibling) cloud, so there’re almost certainly an aunt/uncle/niece/nephew, with a very slim chance that they’re half siblings.
Example 3: Two people at GEDmatch share 1803 cM across 42 segments (the yellow ❸). This one’s a bit more iffy. It’s in a dense part of the red (half sibling) cloud but still well within the black (avuncular) cloud. If this were an unknown parentage case, I would acknowledge to the match that both are possibilities, but that we should explore a half-sibling connection first.
In the three examples, I emphasized that the matches were compared at GEDmatch, because Dr Millard’s simulations were based on the parameters used there. That means that if your data came directly from one of the DNA companies, you should use the chart with caution.
Some considerations:
- AncestryDNA’s phasing algorithm both reduces the total amount of shared DNA and often artificially breaks up continuous blocks of DNA. For example, comparing my two children to their three grandparents who have tested at AncestryDNA, GEDmatch reports an average of 58 more centimorgans and 9.1 fewer segments (about 26% fewer).
- Family Tree DNA includes segments down to 1 cM, while GEDmatch’s threshold is 7 cM. Data from Family Tree DNA must be manually tabulated to include only segments that are 7 cM and larger before using the chart.
- 23andMe includes segments down to 5 cM, so their values should also be adjusted by manually excluding the segments less than 7 cM.
- If you come from an endogamous population, you may share more segments with your relatives than expected, because you are actually related to them more than one way. The additional connections may be numerous and too distant to trace. In this case, the “extra” segments will generally be quite small (under 10 cM). Hopefully, in the future, segment size can be incorporated into our relationship predictions along with total shared DNA and number of segments.
In a forthcoming post, I will describe how simulations by Dr. Millard helped solve a 50-year-old mystery.
UPDATES
Kitty Cooper has just posted a set of empirical (observed, as opposed to simulated) data from self-reported comparisons that addresses the overlap zone. The distinction among groups isn’t as clean-cut with the real-life results. You can read her post here. The difference between Kitty’s data and Andrew’s simulations seem to be attributable to whether the connection is on the maternal or the paternal side. Paternal half-siblings and aunts/uncles/nieces/nephews can sometimes be distinguished from one another, whereas those on the maternal side cannot.
29 Aug 2017 — This post was updated to demonstrate how to get the total cM and number of segments from GEDmatch and to better credit the figures that came from other sources.
1 Sep 2017 — This post was updated to add the second “colors reversed” version of the simulations graph and to add a link to Kitty Cooper’s blog.
8 Oct 2017 — This post was updated with a notice that the distinction between half-sibling and avuncular relationships only holds for paternal relatives. Additional simulations by Andrew Millard indicate that maternal half-siblings and niblings are indistinguishable based on total cM and the number of segments.
~~~~~~~~~~












Hello
Even after reading all you have given I am clueless. I and a first cousin match two sisters and their cousin the two cousins and their cousin all match on chromosome 13 and no other chromosome and if I bring in my first cousin she also matches on chromosome 13 and many other chromosome places. Why do these unknown cousins all show up only on the 13th chromosome and no other?
Thank you for your time and help
Sounds like you’re looking at distant cousins if they only match you on one segment. The best you can do at this point is to narrow the shared ancestor down to the line you share with your first cousin and the line the sisters share with their cousin.
I have an estimated 1st-2nd cousin match at ancestry. I have communicated with this person and her maternal great-grandmother was my maternal grandmother. We share a total of 997cMs on 38 segments. I really wish that ancestry had a chromosome browser, so I could see how long the segment or segments are in chr 23 aka “x”. Another variable in my situation is that my birth father & birth mother were paternal 2nd cousins, so maybe that explains the dna matching at 997cMs on 38 segments, which is more like a 1st cousin then 1st cousin 1x removed.
I would look for another connection to this match besides the 1C1R one that you know about.
She is a shared match to several people and the top (3) are 325cms on 14 segments, 218cMs on 12 segments and 175cMs on 11 segments. The top two of these two matches are also descendants of the same “Moreman” line as my maternal grandmother. I have asked my top 1C1R match to upload her raw data to either http://www.gedmatch.com or to http://www.familytreedna.com so we can compare using their chromosome browsers but she is not responding.
Those are all very good matches. Uploading to GEDmatch and FTDNA might help you find more shared matches, but a chromosome browser isn’t likely to tell you much for matches this close.
I have also communicated with my 2nd highest match, 325cMs on 14 segments, and we are definitely 2nd cousins, related on the same “Moreman” line as my closest match, 997cMs on 38 segments. Our maternal grandmothers were sisters, so we share the same maternal great-grandmother & great-grandfather. I was thinking that if I could see how long the “x” chromosome segment or segments are for my closest match, then maybe that would help me to confirm 1st cousin 1x removed or possibly a different relation?
Need to relabel Grandpa in pink to Grandma.
If you mean the graphic right below the text that reads “The grandmother’s two chromosomes recombine into the uncle like this:”, they’re both grandma’s chromosomes. Remember that she had two copies of chromosome 1 (one from her mom, one from her dad), and those two copies recombined to make the ones that she passed on to her children.
I love the # segments, IBD half distribution graph, although I don’t know why it’s IBD “half” ?!?!?
I gather that it originated from 23 and me ?!?!?
Blaine Bettingers recent [ circa Aug 2017 ] shared cM project chart gives excellent details on cM values, ie low, average, and high, but the distribution chart illustrates the different ranges much better. A picture means 1000 words !!!
The dark green seems to overlap the light blue, so maybe several separate graphs to illustrate the distribution for each scenario might be an improvement for future revisions.
Well done !!!
The graph you are referring to came from a scientific paper published by 23andMe scientists in 2012.
Jesus. Actually, segments of at least 3 cM that do not overlap any segment of at least 1 cM are valuable.
Why on earth are you telling people to discard matches below 7cm?
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3350316/
Because multiple independent comparisons have shown that the smaller the segment, the more likely it is to be a false positive. The tipping point, where an unphased segment has a 50-50 chance of being a real match, is about 7 cM. Even the paper you linked says “the false positive rate can be high for shorter segments.” You may wish to refer to the statistics on the ISOGG wiki.
https://isogg.org/wiki/Identical_by_descent#False_positive_matches
Another problem, which is also addressed in the paper you linked, is that small segments may be IBD in a technical sense (meaning they’re not false positives) but not have any genealogical utility. Put another way, the segments are shared because they are frequent in a given population, not because they were inherited from a recent common ancestor. The shared ancestor who contributed the segment may have lived hundreds of generations ago, and the two people in question may be related dozens of different ways. Thus, the segment has no genealogical use.
On my Ancestry DNA there is someone that is showing 1,845 cM on 62 DNA segments.
This person appears to be a very close relative that I do not know. Can you explain to me how close they may be?
I recommend plugging the cM amount into this tool. It will highlight the possible relationships and tell you which is most likely.
https://dnapainter.com/tools/sharedcmv4
In your case, the match is almost certainly a half sibling, full aunt/uncle/niece/nephew, or grandparent/grandchild.
Two questions:
Can you explain what simulation means in this context?
Why would this only apply to avuncular relationships?
A computer program was used to artificially generate thousands of DNA matches between pairs of relatives. The program lets you quickly collect and analyze lots of data where you know for sure what the relationship is. You can do this for any relationship, but the patterns aren’t necessarily informative for relationships past the avuncular level.
I came across a close match on my Ancestry, her and I are 9 yrs apart. This is our match, 26% shared DNA: 1,796 cM across 36 segments. She has an older brother that’s 2 yrs older than her and a couple of younger siblings so we have ruled out the niece/aunt out. Could we be half siblings?
If you can rule out grandmother/granddaughter (that’s easy given your ages) and aunt/niece (could you be a full sib to their parent), that leaves half-sibling.
Hi, i hope someone can help me
I have an uncle that i know of as an uncle too, but there are quite a lot of secrets.
He is the older of the family but in his Dna comes a much younger age the same age as my parents what i know is not even near right.
My heritage results
26.4%
1868.6 cM
Shared segments 38
Largest segment 170.2 cM
X-dna 180.1(I use a tool to see the X-dna)
His daughter also shares a lot more than expected for first cousin
14.4%
1021.4 cm
34 segments
largest segment:133.5 Cm
X- dna 182 CM
38 shared segments
Is this possible as even his grand daughter is not sharing so much dna and her grandmother his 4 cousin to her grandfather.
also there isnt much possibility that my “Dad” will have any link with mothers family
Because there is a possibility that they arent my parents i would like to find the true
Also i find very strange that most matches they are identified as 2 cousins to me and 4th cousins to him and vice versa i find that odd
Some tools used points for grandfather others for half sibling or uncle but because there aint a lot of percentage differences leaves me confused
Feel free to comment and help out.
I do not know why the only match of a closer relationship is so high and so difficult to get the most accurate :
The match to your uncle (1869 cM) is perfectly reasonable for that relationship. I see no reason to suspect anything other than that he’s your full uncle.